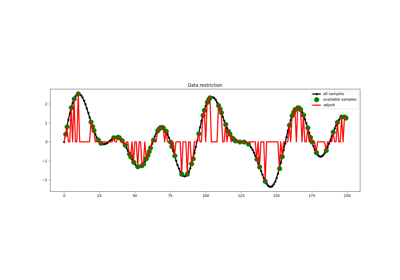
pylops_distributed.Restriction¶
-
class
pylops_distributed.Restriction(M, iava, dims=None, dir=0, inplace=True, compute=(False, False), todask=(False, False), dtype='float64')[source]¶ Restriction (or sampling) operator.
Extract subset of values from input vector at locations
iavain forward mode and place those values at locationsiavain an otherwise zero vector in adjoint mode.Parameters: - M :
int Number of samples in model.
- iava :
listornumpy.ndarray Integer indices of available samples for data selection.
- dims :
list Number of samples for each dimension (
Noneif only one dimension is available)- dir :
int, optional Direction along which restriction is applied.
- inplace :
bool, optional Work inplace (
True) or make a new copy (False). By default, data is a reference to the model (in forward) and model is a reference to the data (in adjoint).- compute :
tuple, optional Compute the outcome of forward and adjoint or simply define the graph and return a
dask.array- todask :
tuple, optional Apply
dask.array.from_arrayto model and data before applying forward and adjoint respectively- dtype :
str, optional Type of elements in input array.
Notes
Refer to
pylops.basicoperators.Restrictionfor implementation details.Attributes: Methods
__init__(M, iava[, dims, dir, inplace, …])Initialize this LinearOperator. adjoint()Hermitian adjoint. apply_columns(cols)Apply subset of columns of operator cond([uselobpcg])Condition number of linear operator. conj()Complex conjugate operator div(y[, niter])Solve the linear problem \(\mathbf{y}=\mathbf{A}\mathbf{x}\). dot(x)Matrix-vector multiplication. eigs([neigs, symmetric, niter, uselobpcg])Most significant eigenvalues of linear operator. matmat(X)Matrix-matrix multiplication. matvec(x)Matrix-vector multiplication. rmatmat(X)Adjoint matrix-matrix multiplication. rmatvec(x)Adjoint Matrix-vector multiplication. todense()Return dense matrix. tosparse()Return sparse matrix. transpose()Transpose this linear operator. - M :